Chi-Square Goodness of Fit
Test Calculator
A free online chi-square goodness of fit test calculator. Enter observed frequencies, compare them to expected counts, and instantly get the chi-square statistic, p-value, Cramér's V effect size, standardized residuals, and an APA-format result string — all in one click.
| Category Name | Observed Count |
|---|
📋 Full Statistical Results
🔢 Observed vs Expected Frequencies
🎨 Colorful Visualizations
🔎 Assumption Checks
📖 Detailed Interpretation of Results
✍️ How to Write Your Results in Research
🎯 Conclusion
🧮 Formulas & Technical Notes
Test Statistic — Pearson's Chi-Square
Degrees of Freedom
Standardized Residual (per cell)
Effect Size — Cramér's V (one-variable case)
Yates' Continuity Correction (only for df = 1)
Technical Notes
- The test statistic follows a χ² distribution with df = k − 1 − m under the null hypothesis.
- The p-value is computed from the upper tail: P(χ² > observed | df).
- If any expected count Eᵢ < 5 in more than 20% of cells, the χ² approximation may be unreliable; consider Fisher's exact test or Monte Carlo simulation.
- Yates' correction is applied only for df = 1 and is conservative — it reduces Type I error but lowers power.
📌 When to Use This Test
This free chi-square goodness of fit test calculator is designed for testing whether observed frequencies in a single categorical variable match a hypothesized distribution. It answers the question: does my sample come from the population I think it does?
Decision Checklist
- You have one categorical variable with two or more mutually exclusive categories
- Your data are frequencies (counts), not means or proportions
- Each observation belongs to exactly one category (mutually exclusive)
- Observations are independent of each other
- Expected count in each category is at least 5 (or 1 in ≤20% of cells)
- Do NOT use if you have two categorical variables → use Chi-Square Test of Independence
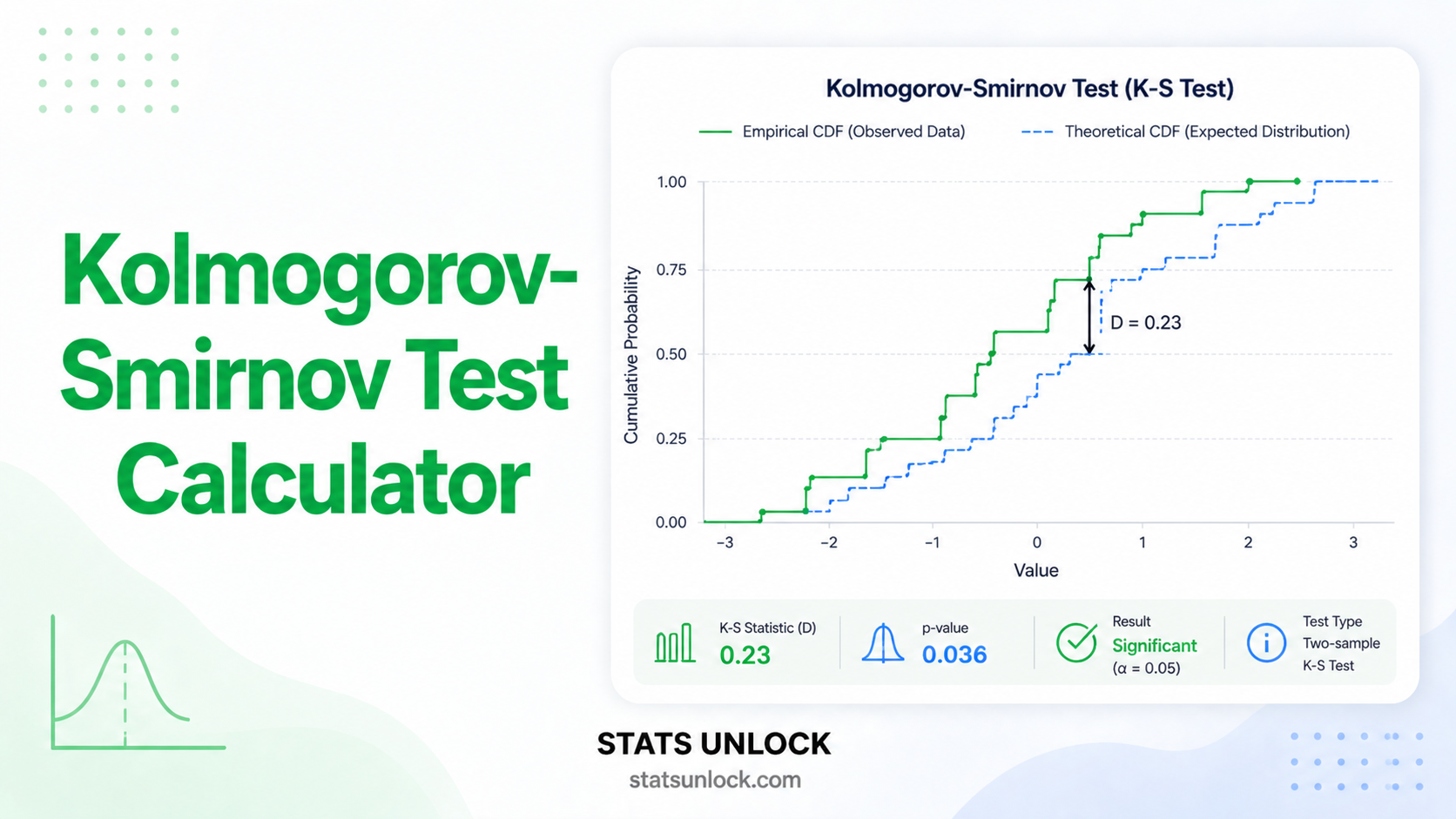
- Do NOT use if your data are continuous → use Kolmogorov-Smirnov or Anderson-Darling
- Do NOT use if observations are paired/repeated on the same subjects → use McNemar's test
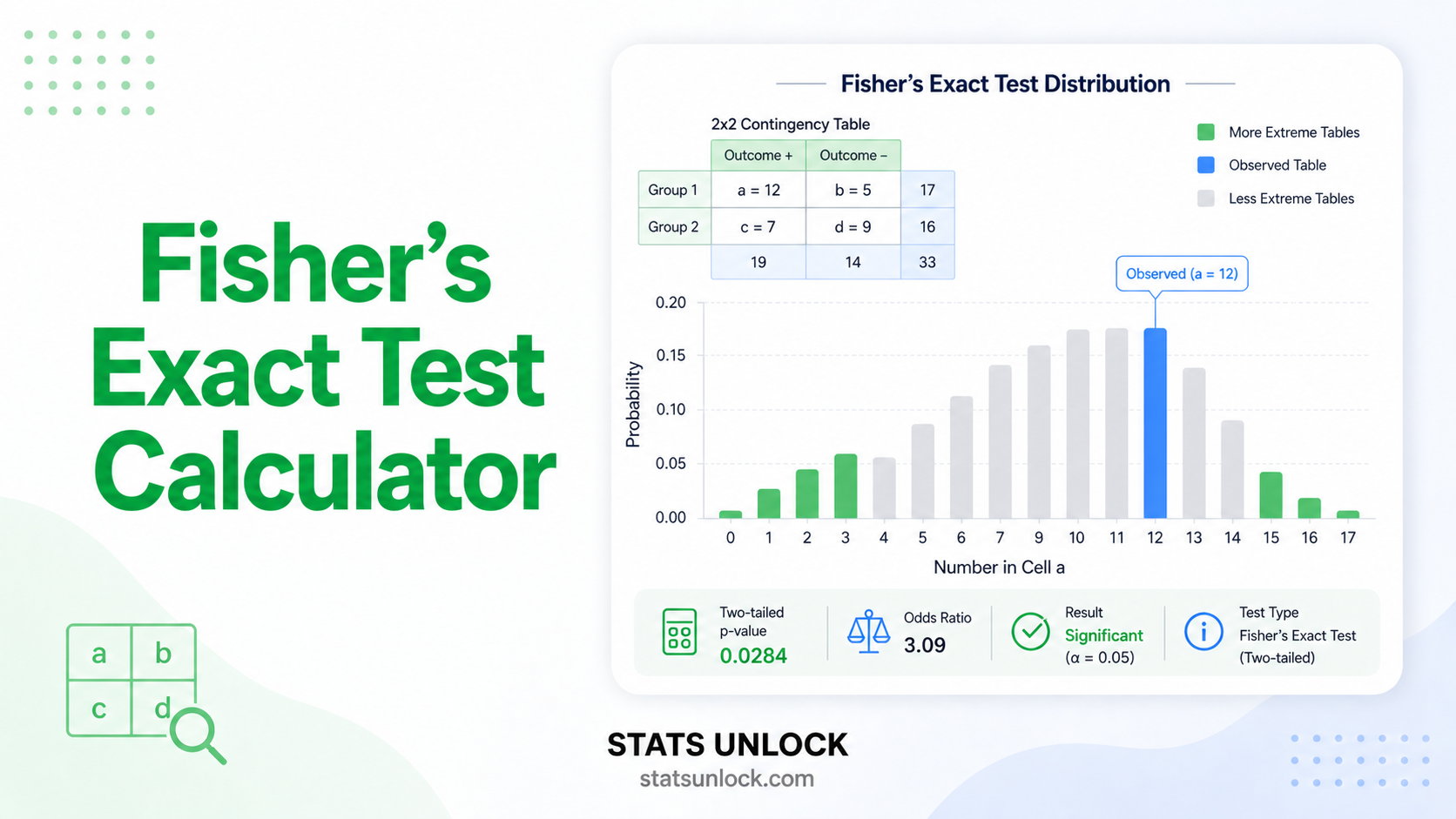
- Do NOT use if expected counts are very small (<1 in any cell) → use Fisher's exact test
Real-World Examples
- Genetics — Mendelian inheritance: A geneticist crosses two heterozygous pea plants and counts 315 round-yellow, 101 wrinkled-yellow, 108 round-green, 32 wrinkled-green offspring. Test whether these counts match the theoretical 9:3:3:1 ratio.
- Quality control — die fairness: A casino auditor rolls a die 600 times and records frequencies for faces 1–6. Test whether the die is fair (each face expected at 1/6 = 100 times).
- Marketing — consumer preference: A company tests five flavors of ice cream. From 500 surveyed customers, observe how many prefer each flavor. Test whether preferences are equally distributed (uniform null) or whether some flavors are preferred.
- Wildlife ecology — habitat selection: A camera-trap study records 240 detections of leopards across four habitat types (forest, grassland, scrub, agriculture). Test whether detections deviate from random use proportional to habitat availability.
- Education — multiple choice answer keys: A teacher analyzes 100-item answer keys to test whether correct answers (A, B, C, D) are uniformly distributed, as a check for test-construction bias.
Sample Size Guidance
Minimum recommended total N: at least 5 × k (so each category's expected count ≥ 5). For small effect detection (V ≈ 0.10), aim for N ≥ 200; for medium (V ≈ 0.30), N ≥ 90; for large (V ≈ 0.50), N ≥ 40.
Decision Tree — Choosing the Right Test
One categorical variable → THIS TEST (Chi-Square Goodness of Fit)
→ Small expected counts (<5 in many cells) → Fisher's exact / Monte Carlo
Two categorical variables → Chi-Square Test of Independence
→ 2×2 small N → Fisher's exact test
→ Paired binary outcomes → McNemar's test
Continuous data → Normality? → Shapiro-Wilk / Anderson-Darling / Kolmogorov-Smirnov
📘 How to Use This Chi-Square Goodness of Fit Calculator — Step-by-Step
Enter Your Data
Choose one of three input methods: type/paste comma-separated counts (default), upload a CSV/Excel file and pick which column holds the category names and which holds the observed counts, or use the manual entry table. Every method results in the same data being analyzed.
Choose a Sample Dataset (Optional)
Five built-in datasets cover the most common use cases: dice fairness, Mendelian 9:3:3:1 inheritance, customer preference, Likert survey distribution, and wildlife habitat selection. Sample 1 (Dice Rolls) loads by default.
Set Category Names and Group Name
Edit the comma-separated category names — these label every table, axis, and chart. Edit the group name to label your dataset (e.g., "Dice rolls", "Pea plants", "Leopard detections") — it shows up in the results, exports, and APA write-up.
Choose Your Expected Frequency Model
Three options: Uniform (equal counts across categories — the default for fairness tests), Custom proportions (e.g., 9:3:3:1 for Mendelian, will be normalized automatically), or Custom counts (enter expected frequencies directly).
Configure the Test
Choose α (default 0.05). If you estimated parameters from your data (e.g., the mean for fitting a Poisson distribution), increase the df reduction. Yates' continuity correction is only for df = 1 cases.
Click "Run Chi-Square Goodness of Fit Test"
Calculation is instant. Results, four colorful visualizations, residuals table, and assumption checks appear below.
Read the Summary Cards
Four colour-coded cards show χ², df, p-value, and Cramér's V. Green = significant at α; amber = non-significant. The cards are the at-a-glance verdict.
Inspect the Visualizations
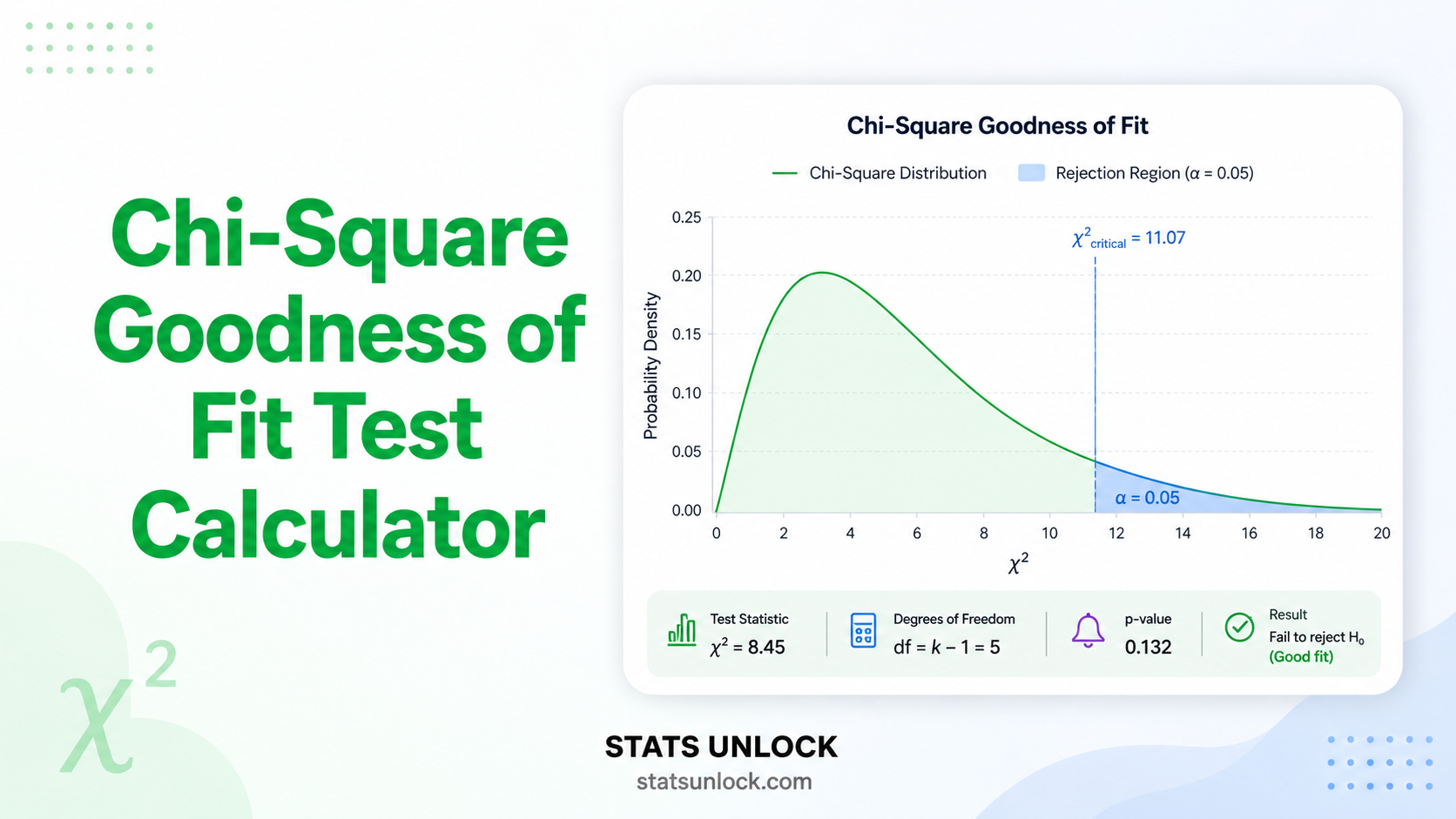
Chart 1 (bars) compares observed vs expected counts. Chart 2 (residuals) shows which categories drive significance — bars beyond ±1.96 are flagged. Chart 3 (donut) shows each category's % contribution to χ². Chart 4 (curve) plots the χ² distribution with the critical region shaded.
Check Assumptions
Three pass/warn/fail badges report (a) sample size adequacy, (b) expected count rule (≥ 5 per cell), and (c) independence. Always satisfy all three before reporting your result.
Export Your Results
Click "Download Doc" for a plain-text .txt report (paste into Word or Google Docs). Click "Download PDF" to print a clean A4-formatted PDF. Use "Copy summary statement" to grab the APA-ready one-liner for your manuscript.
❓ Frequently Asked Questions
Q1. What is the chi-square goodness of fit test and when should I use it?
Q2. What is the difference between chi-square goodness of fit and chi-square test of independence?
Q3. What are the assumptions of the chi-square goodness of fit test?
Q4. How do I interpret a significant result?
Q5. What effect size should I report?
Q6. What are degrees of freedom for chi-square goodness of fit?
Q7. Can I test for a uniform distribution?
Q8. What if my expected frequencies are below 5?
Q9. Is chi-square goodness of fit parametric or non-parametric?
Q10. How do I report the result in APA 7th edition format?
📚 References
The following references support the statistical methods used in this chi-square goodness of fit test calculator, covering p-value interpretation, effect size reporting, and best practices in hypothesis testing with categorical data.
- Pearson, K. (1900). On the criterion that a given system of deviations from the probable in the case of a correlated system of variables is such that it can be reasonably supposed to have arisen from random sampling. Philosophical Magazine, 50(302), 157–175. https://doi.org/10.1080/14786440009463897
- Yates, F. (1934). Contingency tables involving small numbers and the χ² test. Supplement to the Journal of the Royal Statistical Society, 1(2), 217–235. https://doi.org/10.2307/2983604
- Cochran, W. G. (1954). Some methods for strengthening the common χ² tests. Biometrics, 10(4), 417–451. https://doi.org/10.2307/3001616
- Cohen, J. (1988). Statistical power analysis for the behavioral sciences (2nd ed.). Lawrence Erlbaum Associates.
- Cramér, H. (1946). Mathematical methods of statistics. Princeton University Press.
- Agresti, A. (2018). An introduction to categorical data analysis (3rd ed.). John Wiley & Sons.
- Field, A. (2018). Discovering statistics using IBM SPSS statistics (5th ed.). SAGE Publications.
- Howell, D. C. (2013). Statistical methods for psychology (8th ed.). Cengage Learning.
- McHugh, M. L. (2013). The chi-square test of independence. Biochemia Medica, 23(2), 143–149. https://doi.org/10.11613/BM.2013.018
- Sharpe, D. (2015). Chi-square test is statistically significant: Now what? Practical Assessment, Research & Evaluation, 20(8), 1–10. https://doi.org/10.7275/tbfa-x148
- Lakens, D. (2013). Calculating and reporting effect sizes to facilitate cumulative science. Frontiers in Psychology, 4, 863. https://doi.org/10.3389/fpsyg.2013.00863
- American Psychological Association. (2020). Publication manual of the American Psychological Association (7th ed.). APA. https://doi.org/10.1037/0000165-000
- R Core Team. (2024). R: A language and environment for statistical computing. R Foundation for Statistical Computing. https://www.R-project.org/
- NIST/SEMATECH. (2013). e-Handbook of statistical methods — Chi-square goodness of fit test. https://www.itl.nist.gov/div898/handbook/eda/section3/eda35f.htm
- Virtanen, P., Gommers, R., Oliphant, T. E., et al. (2020). SciPy 1.0: Fundamental algorithms for scientific computing in Python. Nature Methods, 17, 261–272. https://doi.org/10.1038/s41592-020-0772-5