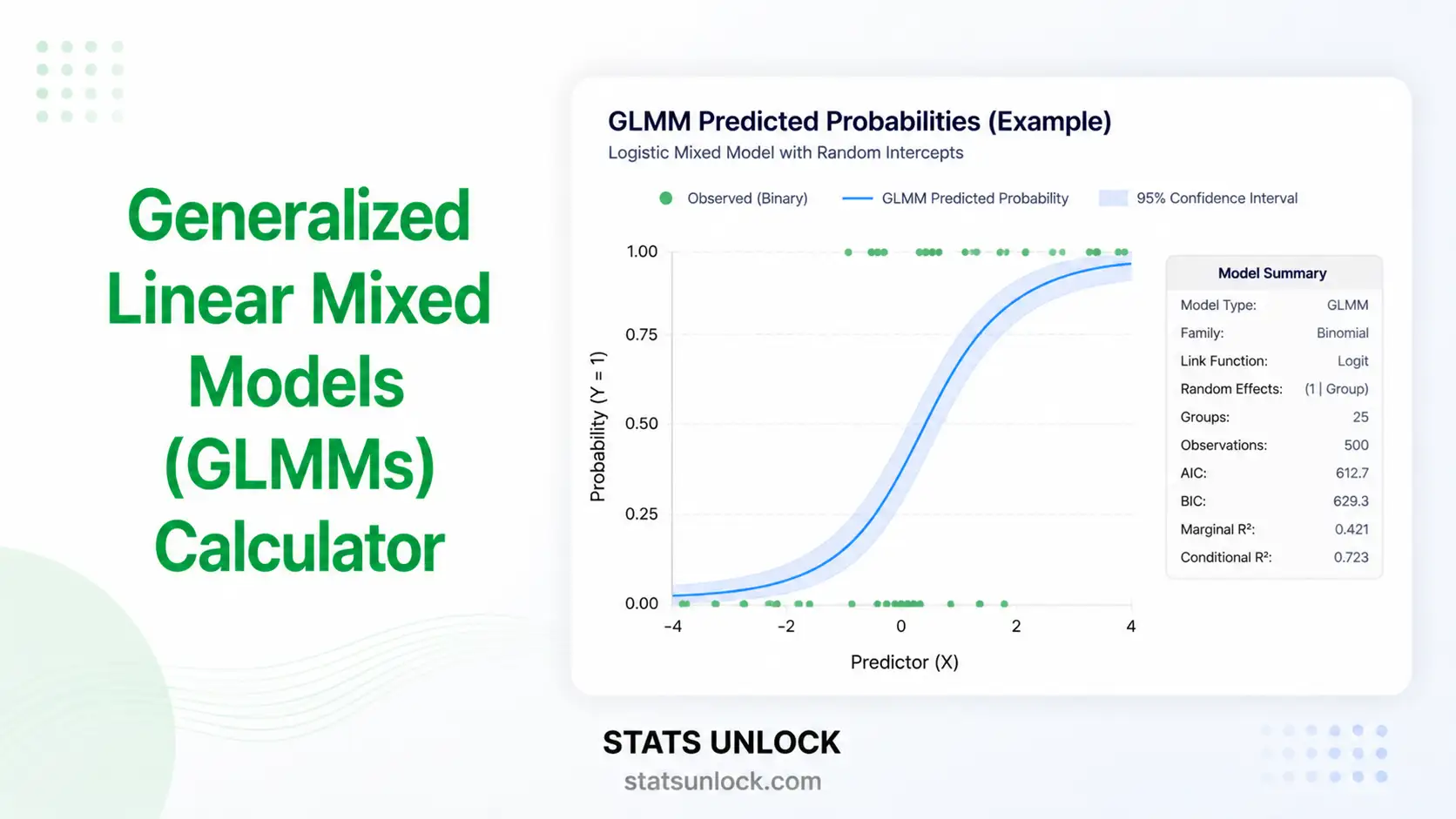
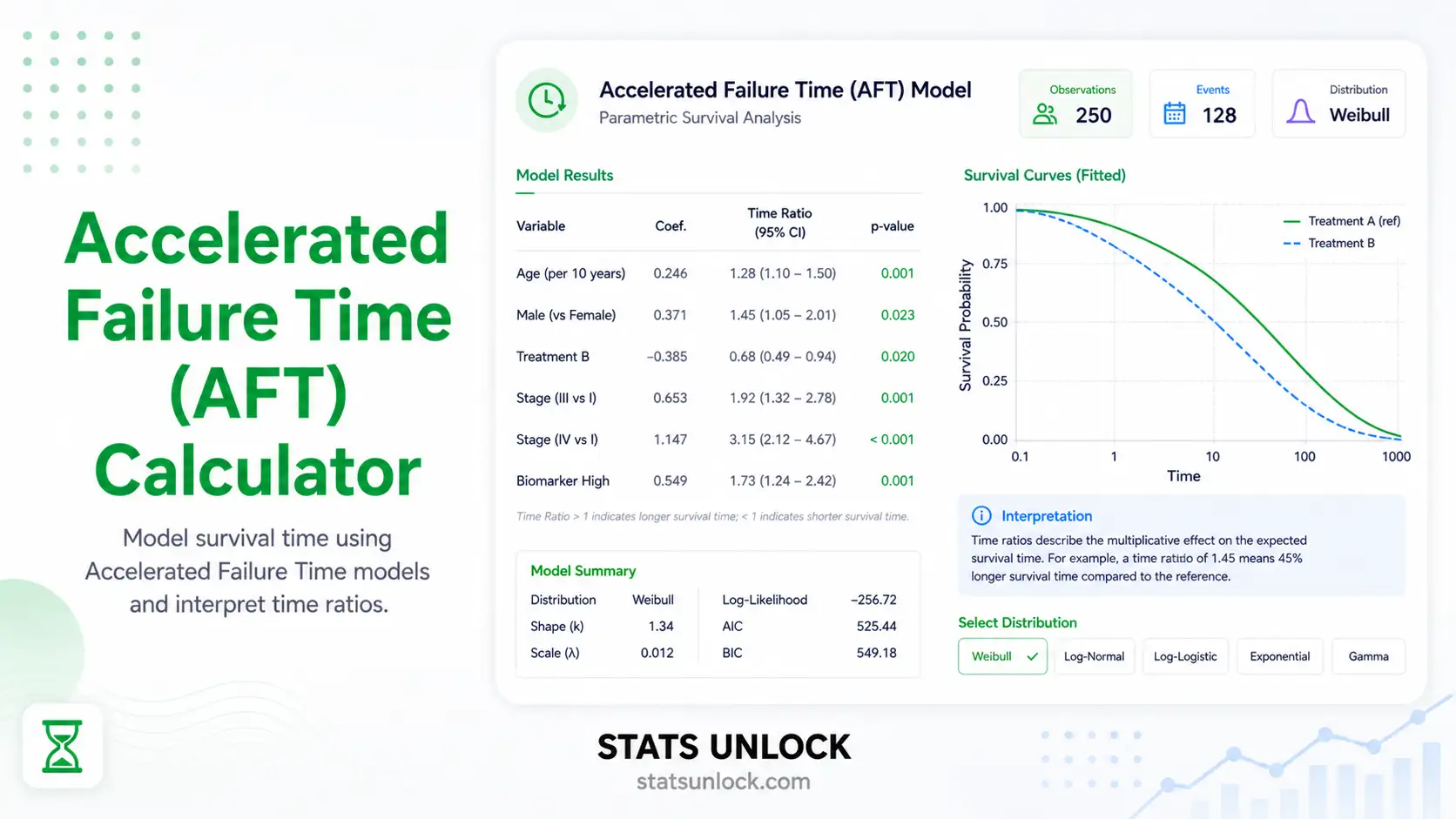
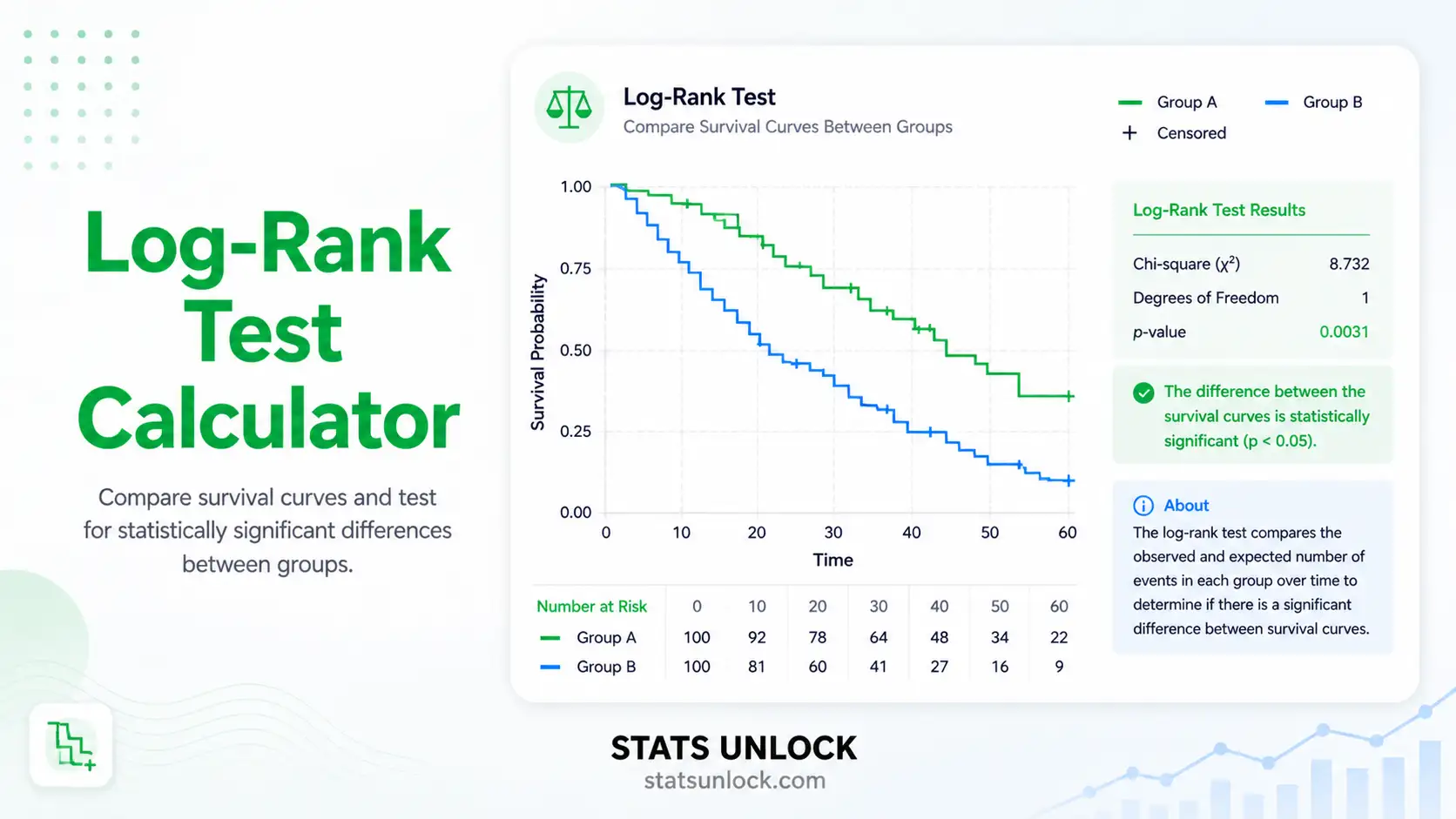
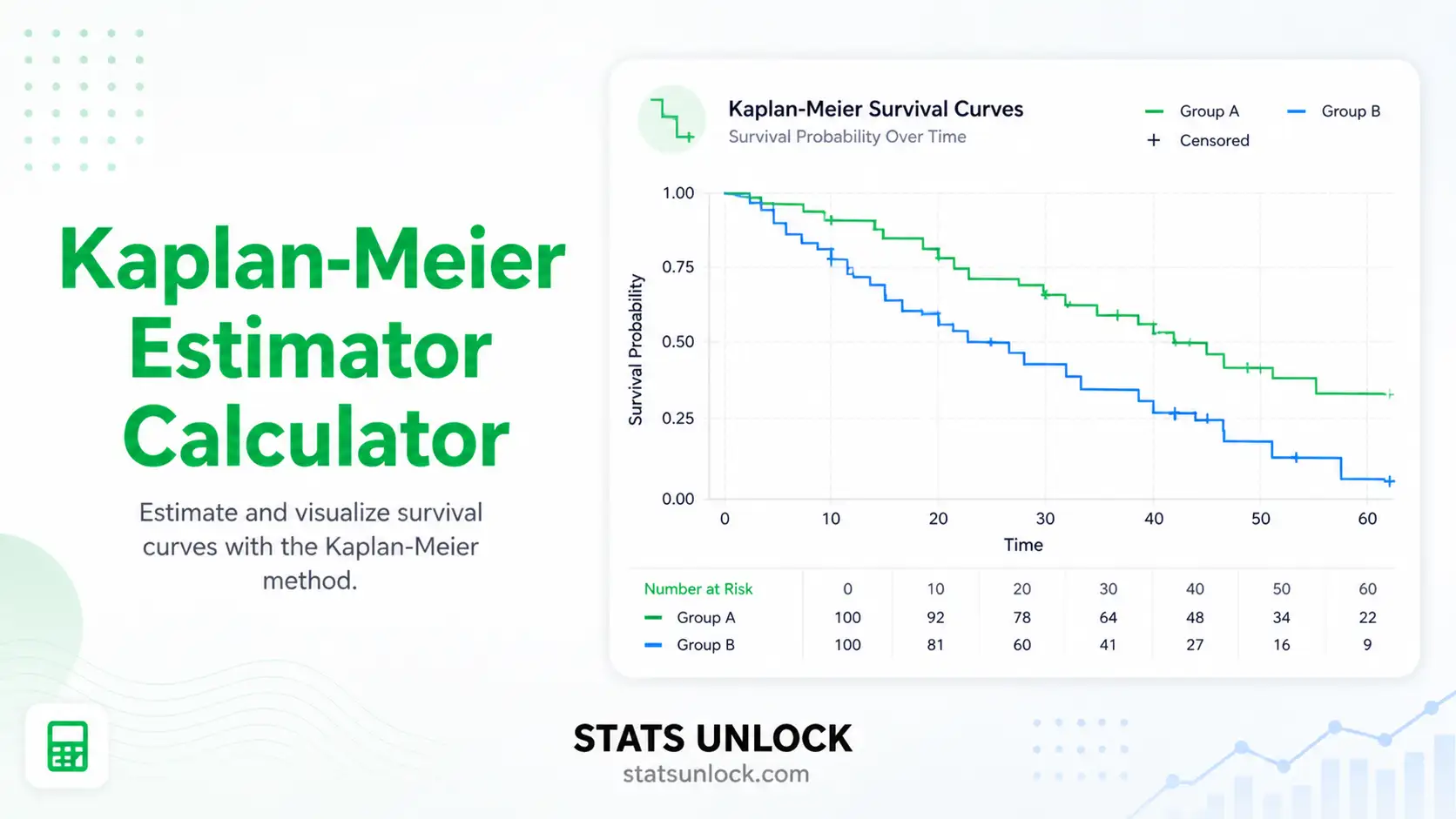
Generalized Additive Mixed Models (GAMM & GLMM) Calculator
Run a free generalized additive mixed model online. Estimate smooth-term effects, random-intercept variance, ICC, AIC, BIC, and adjusted R² across nested or longitudinal data — with APA-format results and four publication-ready visualization plots.
📥 Input Data
Enter each cluster (group / subject / site) as one comma-separated row of numeric responses (Y values). The predictor (X) is auto-generated as the position index 1, 2, 3, … per cluster (use the X-axis option below to relabel). You can rename each group and add or remove clusters as needed.
📂 Upload CSV or Excel file
Accepted: .csv, .txt, .xlsx, .xls. Click columns that should each become a cluster — selected columns will be loaded as separate clusters.
Add a row, type values one per cell, and click Use Manual Data. Each row becomes a separate cluster.
📈 GAMM Results
Detailed statistics
Visualization plots
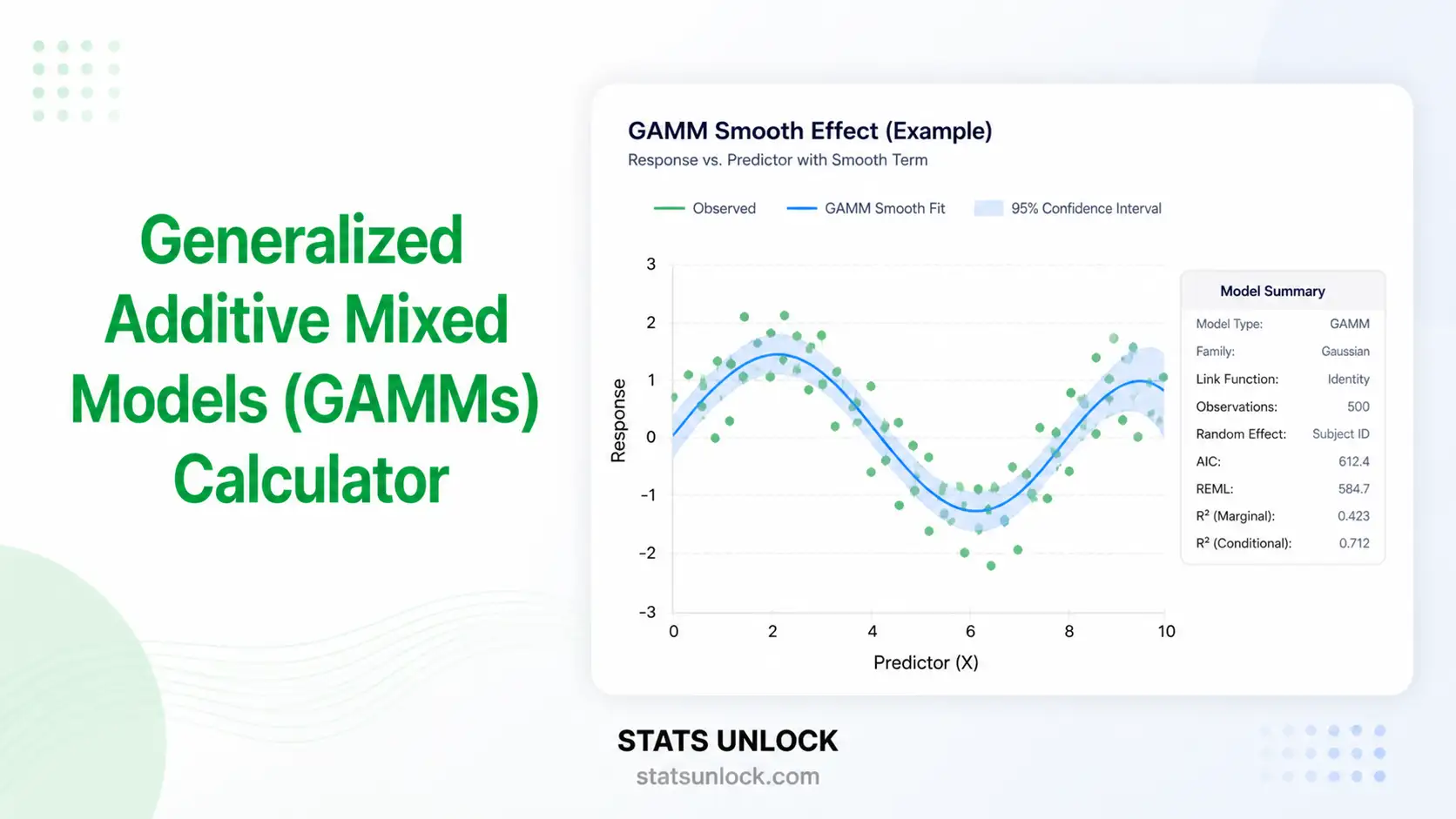
📈 Fitted smooth + raw data
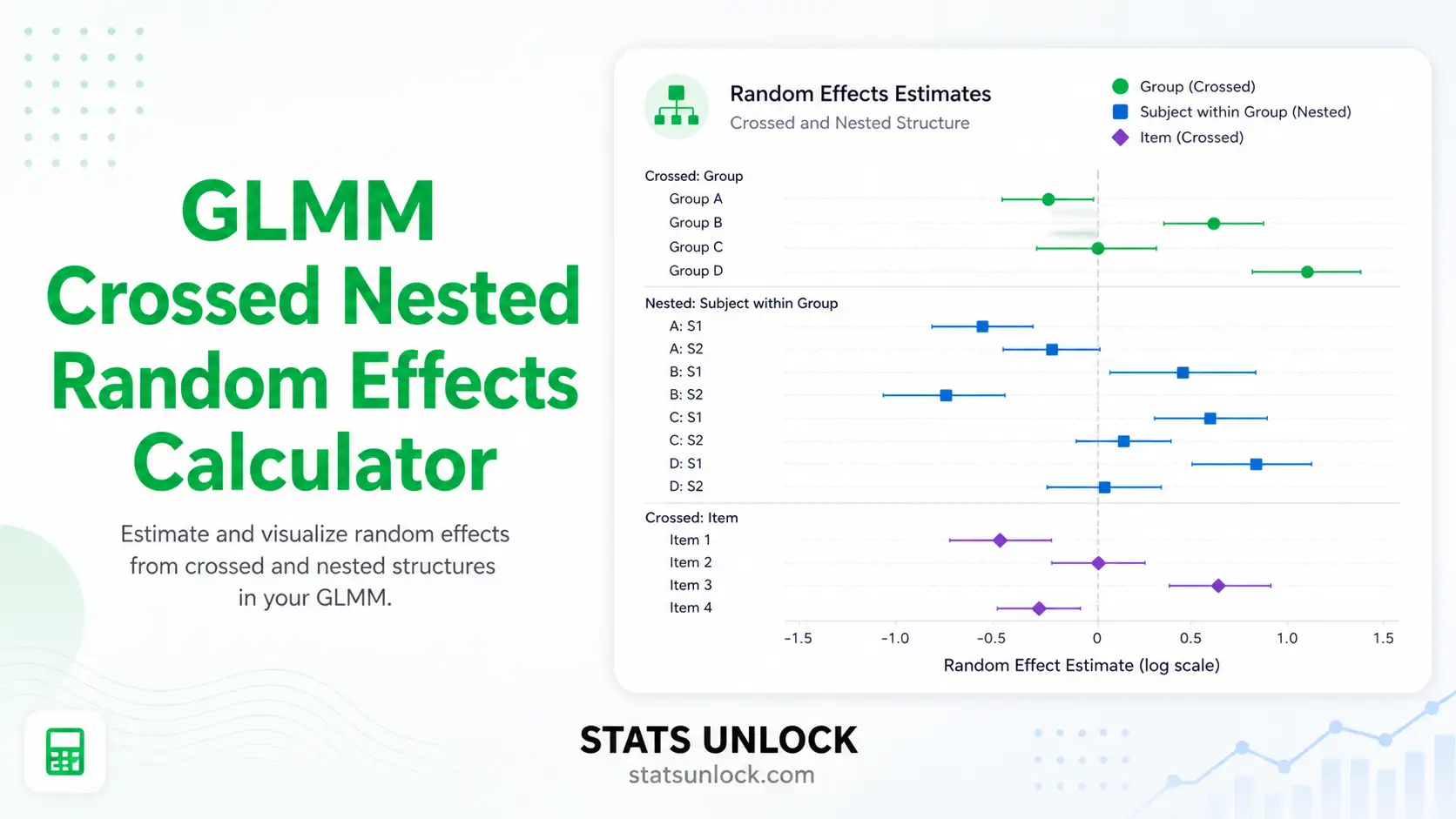
📊 Random effects (cluster intercepts)
🔬 Residual diagnostics
🎯 Q–Q plot of residuals
📐 Technical Notes & Formulas Used
A. Generalized Additive Mixed Model (GAMM) — model equation
y_ij = outcome for observation i in cluster jβ₀ = global intercept (fixed effect)f(x_ij) = smooth (spline) function of predictor x — sum of basis functionsu_j ~ N(0, σ²_u) = random intercept for cluster jε_ij ~ N(0, σ²_ε) = residual error
B. Penalised regression spline (smooth term)
B_k(x) = k-th cubic basis function (truncated power / B-spline)K = basis dimension (user-set, see k in this tool)λ = smoothing parameter chosen by REML / GCVEDF = effective degrees of freedom = trace(F) where F is the influence matrix
C. Variance components (random-effect & ICC)
σ²_u = between-cluster variance (group SD squared)σ²_ε = within-cluster residual varianceICC = intraclass correlation coefficient (0–1); higher → stronger clustering
D. Smooth-term significance test (approximate F)
RSS_null = residual sum of squares with intercept + random effect onlyRSS_full = RSS with smooth term includeddf_resid = n − EDF_smooth − number of clusters
E. Model fit indices
R²_marginal = variance explained by fixed effects only (Nakagawa & Schielzeth, 2013)R²_conditional = variance explained by fixed + random effectslogL = log-likelihood at REML estimates
F. Assumptions of GAMM / GLMM
- Random effects are normally distributed: u_j ~ N(0, σ²_u)
- Residuals are normally distributed and homoscedastic (for Gaussian GAMM)
- Observations within clusters are exchangeable (no unmodelled time dependence beyond the smooth)
- Smooth basis dimension k is large enough (rule: EDF should be well below k)
- At least 5–10 clusters for the random effect to be identifiable
🎯 When to Use GAMM & GLMM Mixed Models
This free generalized additive mixed models calculator is designed for researchers analysing clustered, longitudinal, or repeated-measures data where the predictor's effect is expected to be non-linear. The smooth term captures curvature; the random effect captures cluster correlation. Together they replace the assumptions of plain OLS and standard GLM that fail under both.
Decision checklist
✓ You have a continuous predictor X with possibly non-linear effect
✓ Observations are nested (subjects, schools, sites, plots) — at least 5 clusters
✓ You expect within-cluster correlation (repeated measures, longitudinal data)
✗ Do NOT use if effect of X is clearly linear → use plain GLMM / LMM
✗ Do NOT use if no clustering structure → use GAM (no random effect)
✗ Do NOT use with fewer than 5 clusters → treat group as fixed factor (ANCOVA)
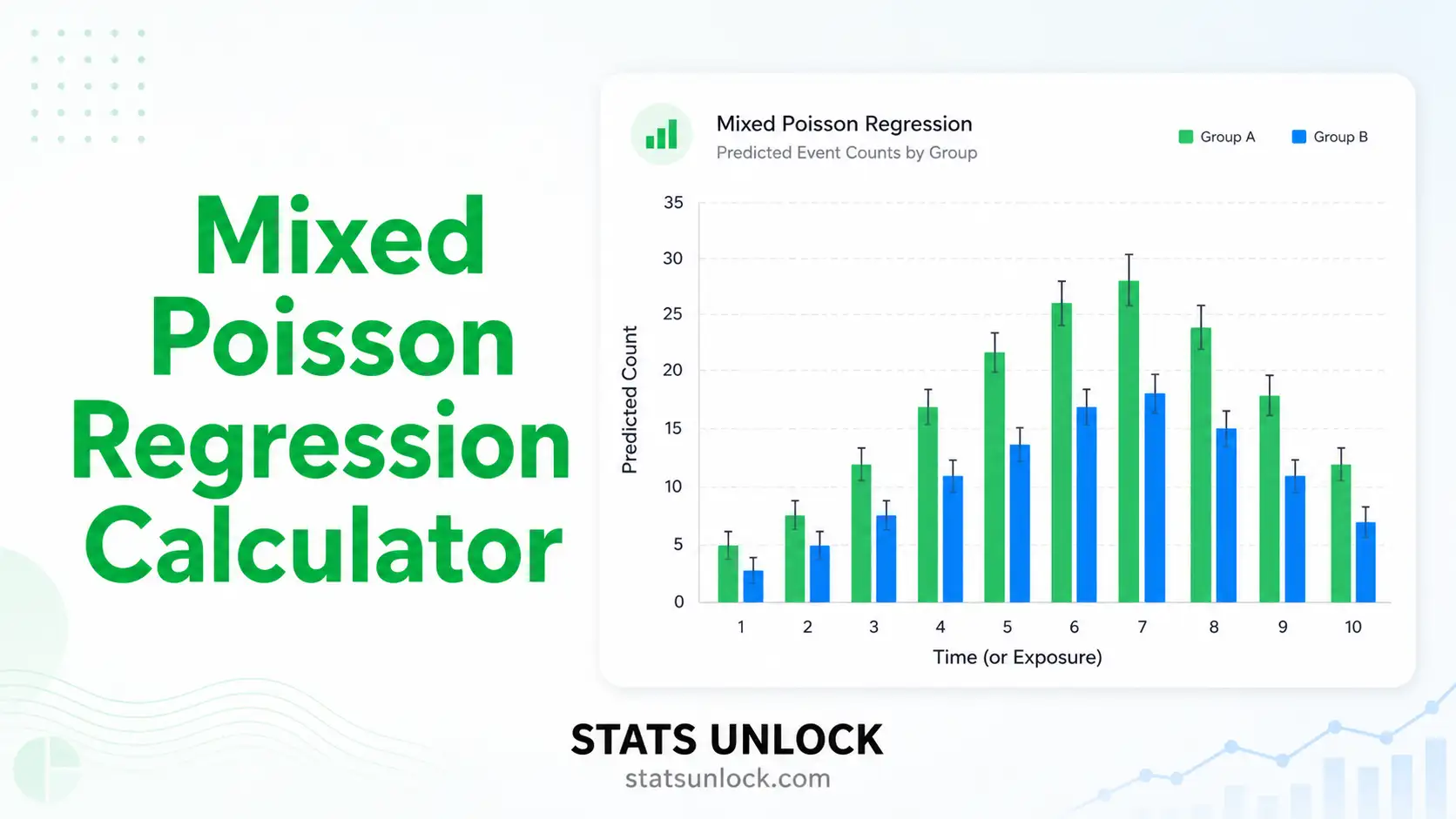
✗ Do NOT use if outcome is binary or count → switch family to binomial / Poisson GAMM
Real-world examples
- Ecology — Modelling plant growth (height) as a non-linear function of light intensity across multiple greenhouses. Greenhouses are random; light effect is smooth.
- Education — Reading test scores over time across multiple schools. Time has a curve (early gain, plateau); schools are random clusters.
- Medicine — Heart-rate response to drug dose across patients in different hospitals. Dose-response is non-linear; hospitals are random.
- Climate science — Forest CO₂ flux as a smooth function of temperature, with random site effects across different sites.
- Animal behaviour — Bird song rate as a smooth function of hour-of-day with random territory effects.
Sample-size guidance
Per cluster: at least 5–10 observations per cluster. Number of clusters: at least 5–10 (Bolker et al., 2009 recommend ≥ 5). Total n: aim for ≥ 10 × k (basis dimension) per smooth term.
Decision tree — which model do I run?
↘ → nested data → LMM (linear mixed model)
↘ non-linear effect → no clustering → GAM
→ nested data → GAMM ← (this tool)
Binary / Count Y → linear, no nesting → GLM
→ linear, nested → GLMM
→ non-linear, nested → GAMM (binomial/Poisson family)
📘 How to Use This GAMM Mixed Models Calculator (Step-by-Step)
- Pick a sample dataset from the dropdown — Plant Growth loads on page open.
- Enter your own data by either typing one comma-separated row per cluster, uploading a CSV/Excel file, or using the manual grid.
- Rename clusters by clicking the group-name field at the top of each row.
- Label your axes using the X-axis name (predictor) and Y-axis name (outcome) inputs.
- Choose smoothness: set the basis dimension k (5 is default). Larger k = more flexible curve.
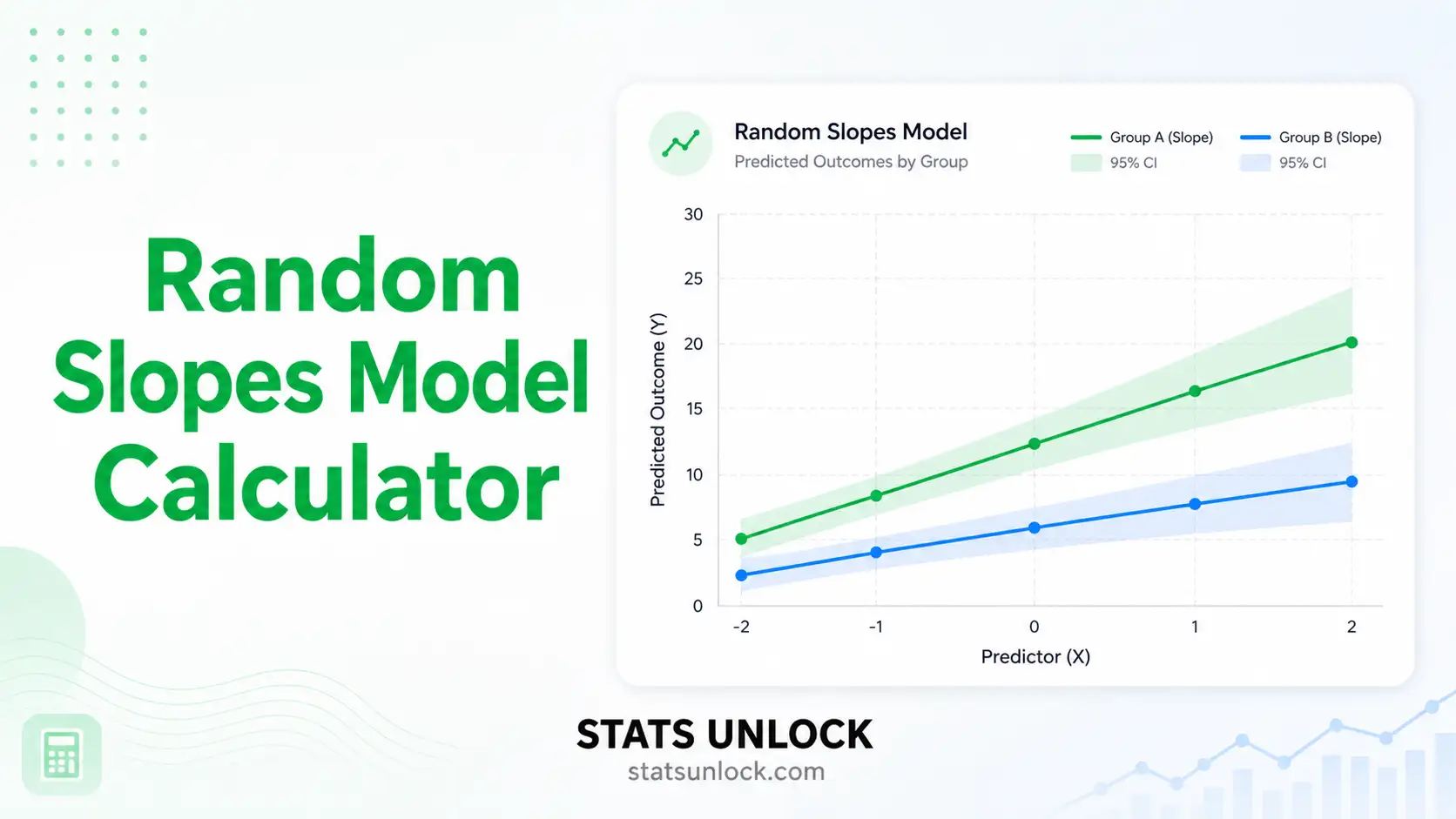
- Set random structure: random intercept only (default) or random intercept + slope.
- Set α to 0.05 (default), 0.01, or 0.10.
- Click "Run GAMM Analysis" — results, four plots, interpretation, and conclusion appear below.
- Read the interpretation — five auto-filled reporting templates (APA, thesis, plain language, abstract, pre-registration) sit ready to copy.
- Export the report as a .txt or save as PDF using the print dialog.
❓ Frequently Asked Questions
What is a generalized additive mixed model (GAMM) and when should I use it?
What is the difference between GAMM and GLMM?
What does the smooth-term p-value mean in a GAMM?
How do I interpret the random-effect variance (group SD) in a mixed model?
What is the ICC (intraclass correlation) in a GAMM or GLMM?
How is AIC used to compare GAMM models?
What sample size do I need for a GAMM or mixed model?
Can I use this calculator for published research?
What does effective degrees of freedom (EDF) mean?
How do I report GAMM results in APA 7th format?
📚 References
The following references support the statistical methods used in this generalized additive mixed models calculator, covering GLMM mixed models, smooth-term significance testing, and best practices in hypothesis testing and reporting.
- Wood, S. N. (2017). Generalized additive models: An introduction with R (2nd ed.). Chapman & Hall/CRC. https://doi.org/10.1201/9781315370279
- Wood, S. N. (2011). Fast stable restricted maximum likelihood and marginal likelihood estimation of semiparametric generalized linear models. Journal of the Royal Statistical Society: Series B, 73(1), 3–36. https://doi.org/10.1111/j.1467-9868.2010.00749.x
- Lin, X., & Zhang, D. (1999). Inference in generalized additive mixed models by using smoothing splines. Journal of the Royal Statistical Society: Series B, 61(2), 381–400. https://doi.org/10.1111/1467-9868.00183
- Pedersen, E. J., Miller, D. L., Simpson, G. L., & Ross, N. (2019). Hierarchical generalized additive models in ecology: An introduction with mgcv. PeerJ, 7, e6876. https://doi.org/10.7717/peerj.6876
- Bolker, B. M., Brooks, M. E., Clark, C. J., Geange, S. W., Poulsen, J. R., Stevens, M. H. H., & White, J.-S. S. (2009). Generalized linear mixed models: A practical guide for ecology and evolution. Trends in Ecology & Evolution, 24(3), 127–135. https://doi.org/10.1016/j.tree.2008.10.008
- Nakagawa, S., & Schielzeth, H. (2013). A general and simple method for obtaining R² from generalized linear mixed-effects models. Methods in Ecology and Evolution, 4(2), 133–142. https://doi.org/10.1111/j.2041-210x.2012.00261.x
- Zuur, A. F., Ieno, E. N., Walker, N. J., Saveliev, A. A., & Smith, G. M. (2009). Mixed effects models and extensions in ecology with R. Springer. https://doi.org/10.1007/978-0-387-87458-6
- Hastie, T., & Tibshirani, R. (1990). Generalized additive models. Chapman & Hall.
- Pinheiro, J. C., & Bates, D. M. (2000). Mixed-effects models in S and S-PLUS. Springer. https://doi.org/10.1007/b98882
- Bates, D., Mächler, M., Bolker, B., & Walker, S. (2015). Fitting linear mixed-effects models using lme4. Journal of Statistical Software, 67(1), 1–48. https://doi.org/10.18637/jss.v067.i01
- Harrison, X. A., Donaldson, L., Correa-Cano, M. E., Evans, J., Fisher, D. N., Goodwin, C. E. D., Robinson, B. S., Hodgson, D. J., & Inger, R. (2018). A brief introduction to mixed effects modelling and multi-model inference in ecology. PeerJ, 6, e4794. https://doi.org/10.7717/peerj.4794
- Simpson, G. L. (2018). Modelling palaeoecological time series using generalised additive models. Frontiers in Ecology and Evolution, 6, 149. https://doi.org/10.3389/fevo.2018.00149
- American Psychological Association. (2020). Publication manual of the American Psychological Association (7th ed.). APA. https://doi.org/10.1037/0000165-000
- R Core Team. (2024). R: A language and environment for statistical computing. R Foundation for Statistical Computing. https://www.R-project.org/
- Wood, S. N. (2024). mgcv: Mixed GAM computation vehicle with automatic smoothness estimation [R package version 1.9-1]. https://cran.r-project.org/package=mgcv