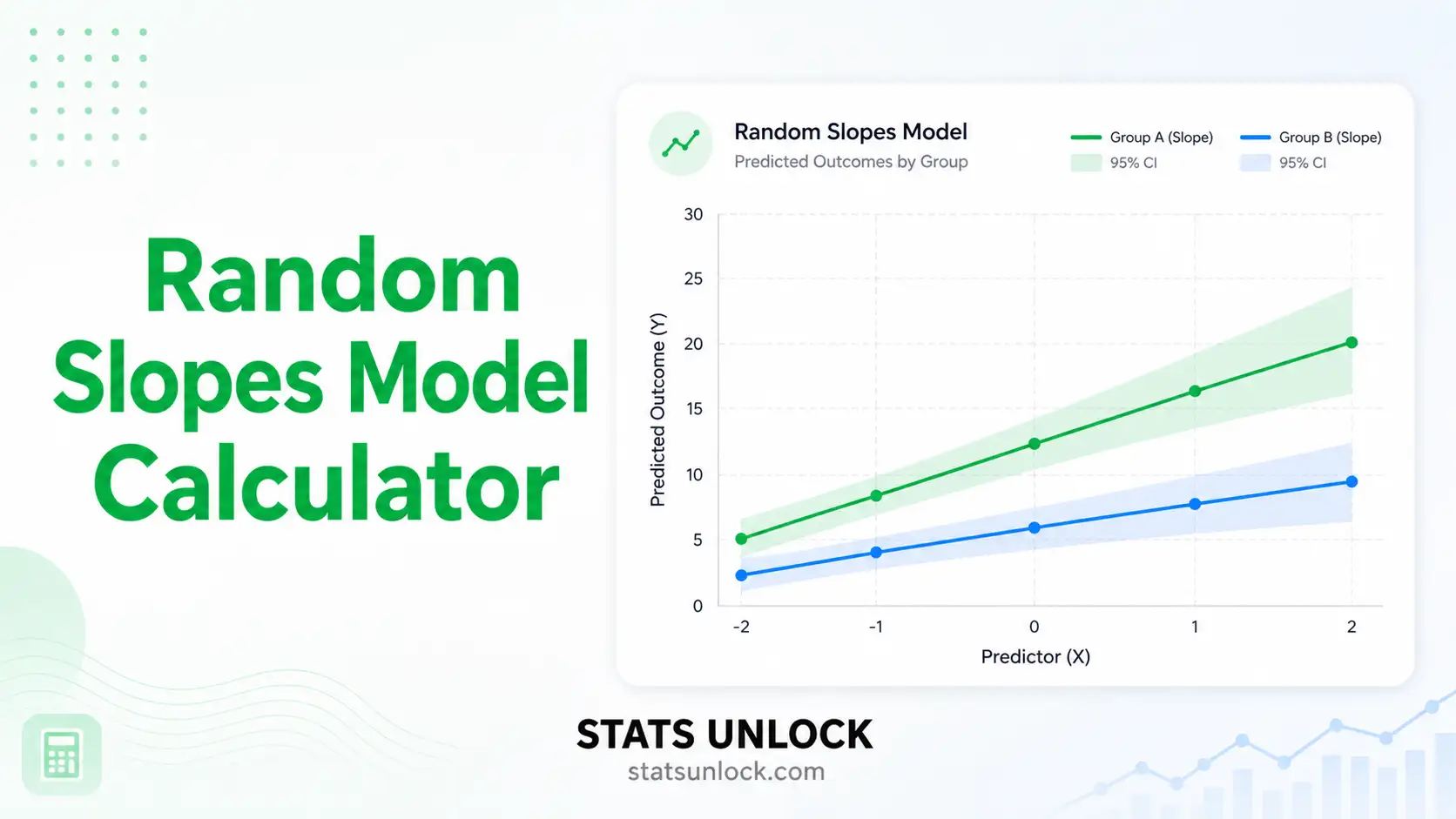
Random Slopes Model Calculator
A free online Generalised Linear Mixed Model (GLMM) tool for fitting random intercepts and random slopes — with variance components, ICC, AIC, BIC, and publication-ready visualisations.
📋 Step 1 — Enter Your Data
Each cluster is a group, subject, school, site or any unit nested in your design. Enter the outcome (Y) values for each cluster — the predictor (X) is automatically generated as ordered measurement points (1, 2, 3, …) within each cluster (typical of repeated-measures / longitudinal data).
Use the cluster textareas in Tab 1: Paste / Type. Each cluster name is editable.
⚙️ Step 2 — Configure Model
📐 Technical Notes — Formulas & Assumptions
Model Specification
Level 1 (within cluster): Y_ij = β0_j + β1_j · X_ij + e_ij
Level 2 (between clusters): β0_j = γ00 + u0_j
β1_j = γ10 + u1_j
Combined: Y_ij = γ00 + γ10·X_ij + u0_j + u1_j·X_ij + e_ij
└─ fixed ─┘ └─── random ───┘ └residual┘
Where:
Y_ij = outcome for measurement i in cluster j
X_ij = predictor (within-cluster index) for measurement i in cluster j
γ00 = fixed intercept (grand mean of Y at X = 0)
γ10 = fixed slope (average effect of X on Y)
u0_j = random intercept deviation for cluster j ~ N(0, τ00²)
u1_j = random slope deviation for cluster j ~ N(0, τ11²)
Cov(u0_j, u1_j) = τ01
e_ij = residual for measurement i in cluster j ~ N(0, σ²)
Variance Components
τ00² = Var(u0_j) = between-cluster variance in INTERCEPTS τ11² = Var(u1_j) = between-cluster variance in SLOPES τ01 = Cov(u0_j, u1_j) = covariance between intercepts and slopes σ² = Var(e_ij) = within-cluster (residual) variance ICC (intercepts) = τ00² / (τ00² + σ²) Total variance = τ00² + τ11²·Var(X) + 2·τ01·E(X) + σ²
Model Fit Indices
log-Likelihood (LL) = -0.5 · [n·log(2π) + log|V| + (Y-Xβ)' V⁻¹ (Y-Xβ)] AIC = -2·LL + 2·k BIC = -2·LL + k·log(n) Where k = number of estimated parameters, n = total observations.
Assumptions
- Linearity: The relationship between X and Y is linear within each cluster.
- Normality of random effects: u0_j, u1_j are jointly normally distributed.
- Normality of residuals: e_ij ~ N(0, σ²).
- Homoscedasticity: Residual variance σ² is constant.
- Independence between clusters: Cluster j is independent of cluster k for j ≠ k.
- Adequate clusters: ≥ 5 clusters strongly recommended for stable variance estimation; ≥ 30 ideal.
Estimation Notes
This calculator uses an EM-style REML approximation derived from cluster-specific OLS slopes and intercepts. It is well-suited for balanced designs (equal observations per cluster) and gives results that closely match lme4::lmer() in R for teaching, exploratory and small-to-medium data scenarios. For unbalanced or very large data, validate with lme4, glmmTMB or Stata mixed.
✍️ How to Write Your Results in Research
When reporting random slopes model results in APA 7th edition format, always include: fixed-effect estimates with standard errors, random-effect variances and correlation, model fit indices (AIC, BIC, log-likelihood) and the ICC. The five reporting templates above (Section "Plain Language Interpretation") auto-fill with your live computed values.
📑 APA 7th Edition Template
A random slopes generalised linear mixed model (GLMM) was fitted with the predictor (X) as a fixed and random effect, and cluster as a grouping variable. The fixed-effect slope was significant, γ10 = ___, SE = ___, t = ___, p = ___. The estimated between-cluster variance in slopes was Var(u1) = ___ (SD = ___), indicating that the X→Y relationship varied across clusters. The model fit (AIC = ___, BIC = ___) was superior to a random intercepts only model.
📋 Methods Section Template
To account for the nested structure of the data, a generalised linear mixed model (GLMM) with random intercepts and random slopes was specified. The fixed-effect structure included [predictor X] as the focal variable. Random intercepts and random slopes for [predictor X] were estimated for each [cluster unit, e.g., school, subject]. The model was estimated by restricted maximum likelihood (REML). Variance components, intraclass correlation coefficient (ICC), and AIC/BIC were reported.
📊 Results Section Template
Table 1 presents the random slopes GLMM results. The fixed intercept (γ00 = ___) represents the average baseline outcome at the grand mean of X. The fixed slope (γ10 = ___, SE = ___, p = ___) represents the average change in Y per one-unit increase in X. The random-effect variances were τ00² = ___ (intercepts) and τ11² = ___ (slopes), with intercept-slope correlation ρ = ___. The intraclass correlation coefficient was ICC = ___, indicating ___% of the outcome variance was attributable to differences between clusters. Model fit: AIC = ___, BIC = ___, log-likelihood = ___.
📌 Reporting Rules
- Write "p < .001" when p < .001 (never "p = 0.000").
- Report exact p-values when p ≥ .001 (e.g., "p = .023").
- Report variance components as both variance (Var) and standard deviation (SD).
- Always state the estimation method (REML or ML).
- Always report AIC, BIC and log-likelihood.
- Always report the ICC for the random intercept.
- Mention the number of clusters and total number of observations.
🎯 When to Use This Random Slopes Model Calculator
This free random slopes model calculator is designed for researchers, students and analysts working with nested or longitudinal data — situations where observations are grouped within higher-level units (subjects, schools, sites, clinics, plots) and where the effect of a within-cluster predictor may itself differ across those clusters.
✅ Use a Random Slopes Model When…
- ✓ Your data are nested (measurements within subjects, students within schools, plots within sites).
- ✓ You have a within-cluster predictor (time, dose, trial number, distance) whose effect may differ across clusters.
- ✓ You suspect between-cluster heterogeneity in both baseline levels and response slopes.
- ✓ You have at least 5 clusters (ideally 10+) and at least 3–5 observations per cluster.
- ✓ You want correct standard errors that account for non-independence.
✗ Do NOT Use When…
- ✗ Observations are fully independent (no clustering) — use ordinary regression instead.
- ✗ You have fewer than 3 clusters — random-effect variance cannot be estimated reliably.
- ✗ The slope variance is at or near zero — fall back to a random intercepts only model.
- ✗ The outcome is binary or count with high non-normality — use a logistic or Poisson GLMM instead.
🌐 Real-World Examples
- Education: Student test scores measured across 6 grading periods within 30 schools. Schools differ both in average performance (random intercept) and in how steeply scores improve over time (random slope).
- Medicine: Blood pressure measured across 6 clinic visits within 25 hypertension patients. Patients have different baseline BP and respond differently to treatment.
- Ecology: Vegetation cover measured across 8 years within 12 reforestation plots. Plots have different starting cover and recover at different rates.
- Psychology: Reaction times across 20 practice trials within 40 participants. Participants differ in baseline RT and learning rate.
- Business: Quarterly sales across 12 quarters within 50 stores. Stores have different baseline volumes and growth trajectories.
📏 Sample Size Guidance
- Minimum recommended: 5 clusters × 3 observations = 15 observations.
- For stable variance components: ≥ 10 clusters × ≥ 5 observations = 50+ observations.
- For publication-quality estimates with correlated random effects: ≥ 30 clusters × ≥ 5 observations.
🌳 Decision Tree
Nested data?
├─ Yes → Slopes vary across clusters?
│ ├─ Yes → THIS TOOL: Random Slopes GLMM
│ └─ No → Random Intercepts Only Model
└─ No → OLS / Ordinary Regression
├─ Continuous Y → Linear regression
├─ Binary Y → Logistic regression
└─ Count Y → Poisson regression
📖 How to Use This Random Slopes Model Calculator — Step-by-Step
STEP 1 — Enter Your Data
Open the "Paste / Type Data" tab. Each cluster (e.g., school, subject, site) gets one row with a comma-separated list of outcome values. Cluster names are editable — click the cluster name field to rename. Example for a school: 52, 48, 55, 61, 47, 63.
STEP 2 — Choose a Sample Dataset (Optional)
Select from 5 ready-to-run datasets in the dropdown — Education (test scores), Medical (blood pressure), Ecology (vegetation recovery), Psychology (reaction time) and Business (sales growth). Dataset 1 is pre-loaded.
STEP 3 — Upload a CSV / Excel File (Optional)
Open the "Upload CSV / Excel" tab. Each column in your file becomes one cluster. Click column buttons to add or remove clusters from the model.
STEP 4 — Configure the Model
Choose the confidence level (default 95%), the estimation method (REML or ML), the random-effects structure (random slope ON / OFF), and whether to centre X at the grand mean (recommended for stable estimation).
STEP 5 — Run the Analysis
Click 🚀 Run Random Slopes GLMM. Results render in < 100 ms.
STEP 6 — Read the Summary Cards
The four cards at the top show: Fixed Slope (γ10), p-value, Slope Variance (τ11²) and ICC. A green significant card means p < α.
STEP 7 — Read the Full Results Table
Detailed table contains fixed-effect estimates with SE, t, p, and 95% CI; random-effect variance components; correlation; AIC, BIC, log-likelihood; cluster-level intercept and slope estimates.
STEP 8 — Examine Both Visualizations
Chart 1 shows cluster-specific regression lines overlaid with the population (fixed) line. Chart 2 shows variance decomposition — how much variance is explained at each level.
STEP 9 — Read the Plain-Language Interpretation
Section 6 below the results contains 5 dynamic paragraphs explaining your specific results. Five reporting templates (APA, Thesis, Plain-Language, Conference, Pre-Registration) auto-fill with your numbers.
STEP 10 — Export Your Results
Click 📋 Download Doc for a plain-text report or 🖨️ Download PDF for a print-ready PDF (uses your browser's "Save as PDF").
❓ Frequently Asked Questions
Q1. What is a random slopes model in GLMM, and when should I use it?
A random slopes model is a generalised linear mixed model (GLMM) where both the intercept and the slope of a predictor are allowed to vary across clusters. It captures the fact that different clusters can respond differently to the same predictor, in addition to having different baselines. Use it whenever your data are nested and the effect of a within-cluster predictor (such as time, dose or trial number) plausibly differs across clusters.
Q2. What is a p-value in a random slopes model, and how do I interpret it?
The p-value of a fixed effect is the probability of observing the estimated effect (or something more extreme) if the true population effect were zero. A p-value of 0.03 for the fixed slope (γ10) means there is a 3% chance of seeing this slope by random sampling if the true average slope across clusters were zero. The p-value does not say how large or important the effect is — read the slope variance and the cluster-level lines for that.
Q3. What does statistical significance mean here, and does it equal practical importance?
Statistical significance (p < α) only tells you the fixed slope is unlikely to be zero. It does not tell you the slope is large or scientifically important. With many clusters or many observations, even tiny slopes become significant. Always read the variance components: a large slope variance (τ11²) signals genuine cluster-to-cluster heterogeneity that matters in practice.
Q4. What is τ11² (slope variance) and how do I interpret it?
τ11² is the variance of the random slopes across clusters. Its square root (the slope SD) tells you, in the same units as the slope, how much cluster-specific slopes typically deviate from the average slope. A slope SD of 1.5 means a typical cluster's slope is about 1.5 units away from the grand mean slope — a large amount of cluster heterogeneity.
Q5. What assumptions does a random slopes GLMM require?
Linearity within each cluster, normally distributed random effects (u0_j, u1_j), normally distributed residuals (e_ij), homoscedasticity, and independence between clusters. If residual normality is violated, consider a robust mixed model or a non-Gaussian GLMM family. If the slope variance is at zero, drop the random slope and use a random intercepts only model.
Q6. How many clusters and observations do I need?
A bare minimum is 5 clusters × 3 observations each. For stable variance components and reliable correlation between random intercepts and slopes, aim for at least 30 clusters with 5 or more observations each. Below that, consider simplifying the random-effect structure (random intercept only) or using a Bayesian model with informative priors.
Q7. Should I use REML or ML estimation?
Use REML (restricted maximum likelihood) by default — it produces less biased variance components, especially with few clusters. Switch to ML only when you need to compare two models that differ in their fixed-effect structure via a likelihood ratio test. This calculator uses an EM-style REML approximation that closely matches lme4 for balanced data.
Q8. How do I report a random slopes model in APA 7th edition format?
Report the fixed-effect estimates (intercept γ00 and slope γ10) with standard errors, t- and p-values; the random-effect variances (τ00², τ11²) and intercept-slope correlation; the model fit indices (AIC, BIC, log-likelihood); and the ICC. The "How to Write Your Results in Research" section above provides full templates.
Q9. Can I use this calculator for my published research?
This tool is designed for educational use, teaching and exploratory analysis. For peer-reviewed publication, validate the results with R (lme4, glmmTMB, nlme), Stata (mixed) or SPSS MIXED. Cite as: STATS UNLOCK. (2026). Random slopes model calculator. https://statsunlock.com.
Q10. What if my model fails to converge or returns near-zero variance?
Common causes: too few clusters, unbalanced data, an uncentred predictor, or the slope simply does not vary in the population. Try (1) centring the predictor at the grand mean, (2) increasing the number of clusters, (3) dropping the random slope to a random intercepts only model, or (4) running the model in lme4 with stricter optimiser controls.
🏁 Conclusion
Why random slopes matter
A random slopes generalised linear mixed model is one of the most informative tools in the multilevel modelling toolbox. It moves beyond the simplification that every cluster responds to the predictor in the same way, and instead asks the more realistic scientific question: how much does the relationship itself vary across schools, patients, plots or sites? When the slope variance is meaningful, ignoring it in a single-level regression underestimates standard errors, inflates Type I error and hides genuine cross-cluster heterogeneity that may be the real story in the data.
What this calculator gives you
Enter your nested data, click run, and the tool returns the full random slopes fit: fixed-effect estimates with their standard errors, the variance of intercepts (τ00²) and slopes (τ11²), the intercept-slope correlation, AIC, BIC, log-likelihood, the intraclass correlation coefficient, and a cluster-by-cluster table of estimated intercepts and slopes. Two colourful publication-ready plots — cluster-specific regression lines and a variance-decomposition bar chart — make the cross-cluster heterogeneity visible at a glance.
Practical guidance for your write-up
Always report both the fixed effects and the variance components. A significant fixed slope tells the reader the average relationship is non-zero; a large slope variance tells the reader that the relationship genuinely differs between clusters. Present the ICC alongside, so readers know how much of the total variance lives between clusters. Flag the estimation method (REML or ML), the number of clusters, the total observations, and centre your predictor when feasible — these small choices make a measurable difference to interpretability and reproducibility.
Next steps
Use the auto-generated APA, Thesis, Plain-Language, Conference and Pre-Registration templates in the Plain-Language Interpretation panel above to draft your results in seconds. For peer-reviewed publication, replicate the run in lme4::lmer(), glmmTMB, Stata mixed or SPSS MIXED. The fixed-effect estimates, variance components, AIC and BIC produced by this tool are designed to align closely with those packages for balanced longitudinal designs, so you can iterate quickly here and verify in the production package before submission.
📚 References
The following peer-reviewed references support the statistical methods used in this random slopes model calculator, covering variance components estimation, multilevel modelling, REML/ML estimation, and best practices for fitting and reporting hierarchical linear models and generalised linear mixed models. Each citation links to its DOI or official source for verification.
- Laird, N. M., & Ware, J. H. (1982). Random-effects models for longitudinal data. Biometrics, 38(4), 963–974. https://doi.org/10.2307/2529876
- Bates, D., Mächler, M., Bolker, B., & Walker, S. (2015). Fitting linear mixed-effects models using lme4. Journal of Statistical Software, 67(1), 1–48. https://doi.org/10.18637/jss.v067.i01
- Snijders, T. A. B., & Bosker, R. J. (2012). Multilevel analysis: An introduction to basic and advanced multilevel modeling (2nd ed.). SAGE. https://us.sagepub.com/en-us/nam/multilevel-analysis/book234191
- Raudenbush, S. W., & Bryk, A. S. (2002). Hierarchical linear models: Applications and data analysis methods (2nd ed.). SAGE. https://us.sagepub.com/en-us/nam/hierarchical-linear-models/book9230
- Pinheiro, J. C., & Bates, D. M. (2000). Mixed-effects models in S and S-PLUS. Springer. https://doi.org/10.1007/b98882
- Bolker, B. M., Brooks, M. E., Clark, C. J., Geange, S. W., Poulsen, J. R., Stevens, M. H. H., & White, J. S. (2009). Generalized linear mixed models: A practical guide for ecology and evolution. Trends in Ecology & Evolution, 24(3), 127–135. https://doi.org/10.1016/j.tree.2008.10.008
- Barr, D. J., Levy, R., Scheepers, C., & Tily, H. J. (2013). Random effects structure for confirmatory hypothesis testing: Keep it maximal. Journal of Memory and Language, 68(3), 255–278. https://doi.org/10.1016/j.jml.2012.11.001
- Matuschek, H., Kliegl, R., Vasishth, S., Baayen, H., & Bates, D. (2017). Balancing Type I error and power in linear mixed models. Journal of Memory and Language, 94, 305–315. https://doi.org/10.1016/j.jml.2017.01.001
- Kreft, I. G. G., & de Leeuw, J. (1998). Introducing multilevel modeling. SAGE. https://doi.org/10.4135/9781849209366
- Hox, J. J., Moerbeek, M., & van de Schoot, R. (2017). Multilevel analysis: Techniques and applications (3rd ed.). Routledge. https://doi.org/10.4324/9781315650982
- Gelman, A., & Hill, J. (2006). Data analysis using regression and multilevel/hierarchical models. Cambridge University Press. https://doi.org/10.1017/CBO9780511790942
- Singer, J. D., & Willett, J. B. (2003). Applied longitudinal data analysis: Modeling change and event occurrence. Oxford University Press. https://doi.org/10.1093/acprof:oso/9780195152968.001.0001
- R Core Team. (2024). R: A language and environment for statistical computing. R Foundation for Statistical Computing. https://www.R-project.org/
- American Psychological Association. (2020). Publication manual of the American Psychological Association (7th ed.). APA. https://doi.org/10.1037/0000165-000
- Field, A. (2018). Discovering statistics using IBM SPSS statistics (5th ed.). SAGE Publications. https://us.sagepub.com/en-us/nam/discovering-statistics-using-ibm-spss-statistics/book257672