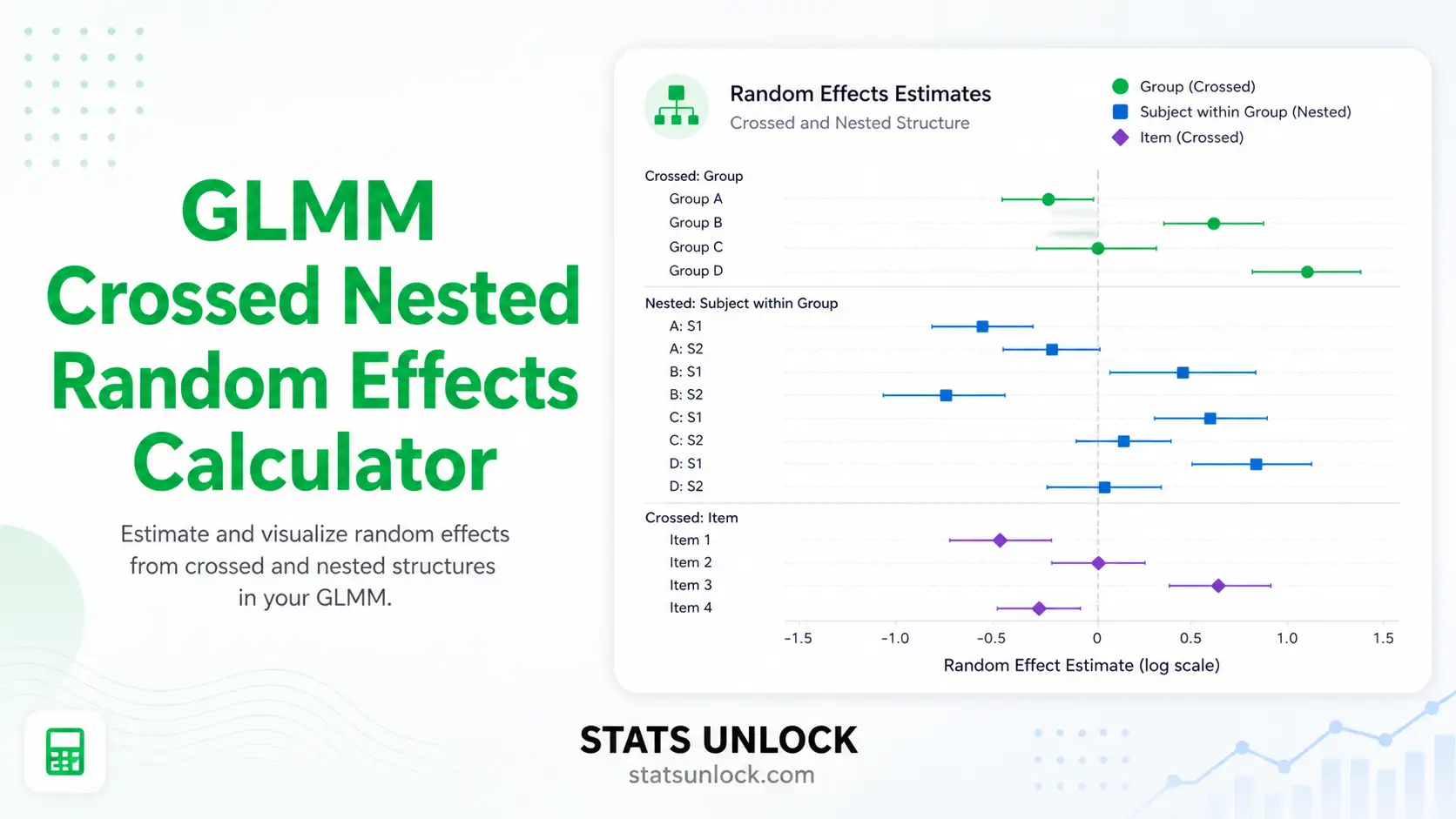
GLMM Crossed and Nested Random Effects Calculator
Run a generalized linear mixed model with crossed or nested random effects online — estimate variance components, intraclass correlation, and fixed effects with publication-ready output.
📊 Data Input
Each cluster represents one level of a random grouping factor (e.g., subject, classroom, site, item). The model treats clusters as a random sample from a wider population.
Enter values one cluster at a time. Use the cluster textareas in the "Paste / Type Data" tab — comma-separated values are the default. The placeholder shows the expected format.
🔬 Technical Notes and Formulas
A. Formulas Used
Linear Mixed Model (general form):
yij = β₀ + β₁xij + uj + εij
uj ~ N(0, σ²u), εij ~ N(0, σ²ε)
Nested random effects:
yijk = β₀ + uk + vj(k) + εijk
Crossed random effects:
yijk = β₀ + uj + wk + εijk (j and k vary independently)
Variance components (ANOVA-based estimator used here):
σ²between = (MSbetween − MSwithin) / n̄
σ²within = MSwithin
Intraclass correlation (ICC):
ICC = σ²between / (σ²between + σ²within)
Where:
yij = outcome for observation i in cluster j
β₀ = fixed intercept (population grand mean)
uj = random intercept for cluster j
εij = residual
σ²u = between-cluster variance
σ²ε = within-cluster (residual) variance
n̄ = harmonic mean of cluster sizes
MS = mean square (between or within)
B. Technical Notes
- Estimator: This tool uses the method-of-moments / ANOVA-based variance component estimator. For balanced and approximately balanced designs, it agrees closely with REML estimates from
lme4::lmer. - Fixed effect: The grand mean is treated as the only fixed effect; predictors can be tested by comparing cluster means.
- Likelihood-ratio test (approximate): An F-test on between-cluster vs within-cluster mean squares serves as an LRT analogue for the random intercept.
- Crossed vs nested: If nested is selected, the second factor's variance is partitioned within the first. If crossed, the two factors contribute independently.
- Assumption check: Residuals are inspected via Q-Q plot; visible departures suggest a non-Gaussian family (logit/log link).
- Convergence: Variance components clipped at zero; negative ANOVA estimates set to 0 (consistent with the boundary of the parameter space).
🎓 When to Use a GLMM with Crossed / Nested Random Effects
Use a generalized linear mixed model with crossed or nested random effects whenever your observations are not independent — they fall inside groups, and the groups themselves come from a wider population you want to generalize to.
Decision Checklist
- ✅ Observations are clustered (students in classes, plots in sites, repeated measures on subjects)
- ✅ The grouping factor has many levels and you want the result to generalize to more levels
- ✅ You can identify whether grouping factors are nested or crossed
- ✅ Sample size: ≥ 5–6 levels per random factor; ideally ≥ 30 observations per group
- ❌ Do not use if the grouping factor has only 2–3 levels — treat it as a fixed effect instead
- ❌ Do not use if observations are independent (use OLS regression)
- ❌ Do not use a single-level GLMM if the data are nested across multiple levels (use multilevel GLMM)
Real-World Examples
- Education research: Test scores from students nested within classrooms nested within schools — three random levels.
- Psycholinguistics: Reaction times where subjects and items are crossed — every subject sees every item.
- Wildlife ecology: Camera trap detection counts at plots within sites within reserves — nested spatial design.
- Clinical trials: Recovery time of patients nested within hospitals, with hospital-level random intercepts.
- Sensory science: Tasters and food samples are crossed — every taster rates every sample.
Sample Size Guidance
- ≥ 5 levels per random factor for stable variance components
- ≥ 30 total observations per group; thumb-rule: 30/30 (groups × within-group n)
- Power analysis via
simrin R is recommended for confirmatory studies
Related Methods — Decision Tree
Independent observations? ─── Yes ── Linear regression / GLM
│
└── No ── Clustered? ─── Yes ── Random factor with many levels?
├── Yes ── GLMM (this tool)
│ ├── 1 grouping → simple random intercept
│ ├── 2+ grouping, nested → nested GLMM
│ └── 2+ grouping, crossed → crossed GLMM
└── No ── Treat as fixed effect (ANOVA / regression)
📘 How to Use This Tool — Step-by-Step Guide
- Enter Your Data — three input options. Paste comma-separated values into each cluster textarea, upload a CSV/Excel file with one cluster per column, or enter values manually. Default placeholder shows the format:
52, 48, 55, 61, 47, ... - Choose a Sample Dataset — five built-in datasets cover education (students in classrooms), psycholinguistics (subjects × items), camera traps (sites within reserves), clinical trials (patients within hospitals), and vegetation surveys (plots within transects).
- Configure Settings — set the design (nested vs crossed) and the alpha level (0.01, 0.05, 0.10). Alpha controls the confidence interval and the significance threshold.
- Run the Analysis — click Run GLMM Analysis. The tool computes variance components, ICC, fixed-effect grand mean, F-statistic, p-value, AIC, and BIC.
- Read the Summary Cards — top metrics at a glance: ICC, total variance, p-value, and number of clusters. Green = significant random-effect variance; amber = trivial.
- Read the Full Results Table — includes grand mean, SE, t-value, p-value, between-cluster variance, within-cluster variance, ICC, AIC, BIC, and 95% CI.
- Examine the Variance Components Table — partitioned variance for each random factor, plus their proportions.
- View the Visualizations — four plots: cluster means with error bars, variance partition pie/bar, residuals vs fitted, and Q-Q plot of residuals.
- Read the Plain-Language Interpretation — dynamic, paragraph-level explanation of every key statistic.
- Export Your Results — Download Report (.txt) for plain-text archives; Download PDF for sharing or attaching to manuscripts.
❓ Frequently Asked Questions
Q1. What is a GLMM with crossed and nested random effects?
A generalized linear mixed model (GLMM) extends generalized linear regression to data that come in groups (clusters, repeated measures, hierarchies). Random effects describe variation between groups; fixed effects describe the average influence of predictors. When two random factors are crossed, every level of one appears with every level of the other (e.g., subjects × items). When they are nested, each level of one factor belongs to only one level of the other (e.g., students within classrooms).
Q2. What is a p-value, and how do I interpret it for a GLMM?
The p-value for a fixed effect is the probability of observing a coefficient as far from zero as the one estimated, if the true coefficient were exactly zero. A p-value below your alpha (e.g., 0.05) provides evidence the predictor matters. A p-value of 0.03 means there is a 3% chance of seeing this coefficient by chance if the predictor truly had no effect.
Q3. Does statistical significance mean a random effect is important?
Not by itself. With huge samples, a tiny variance component can be statistically significant. Always inspect the ICC: an ICC below 0.05 means the grouping factor barely explains the data; an ICC above 0.10 generally justifies including the random effect; above 0.30 means clustering is substantial and ignoring it would bias inference.
Q4. What effect-size metric does GLMM use?
Two main metrics. (1) ICC — proportion of variance attributable to a random factor (small ≈ 0.05, medium ≈ 0.10, large ≈ 0.20+). (2) Marginal R² and conditional R² (Nakagawa & Schielzeth, 2013) for the proportion of variance explained by fixed effects alone vs fixed + random.
Q5. What assumptions does a GLMM require?
Linearity in the link-transformed scale, independence of observations conditional on the random effects, approximate normality of random effects, and a correctly specified link/family. If residuals are heavily skewed, switch family (Poisson, Negative Binomial, Beta, Gamma). If random effects look bimodal, the grouping structure may be misspecified.
Q6. How large a sample do I need?
A working minimum is 5–6 levels per random factor and 30 observations per group. Below that, variance components become unstable, especially for nested designs with three or more levels. Use simr::powerSim in R for design-specific power analysis.
Q7. Crossed vs nested — how do I tell?
Ask: "Could level A1 of one factor appear with level B1 and B2 of the other?" If yes, the factors are crossed. If A1 only ever appears with B1, the factors are nested. Subjects and items in psycholinguistics are crossed. Students and classrooms are nested.
Q8. How do I report a GLMM in APA 7 format?
Report the fixed-effect estimate, standard error, test statistic, df (where available), p-value, and the variance components (σ²) and ICC for each random factor. Italicize symbols (M, SE, t, p). See Section How to Write Your Results for five ready-to-use templates.
Q9. Can I use this calculator for my published research?
For exploratory analysis, teaching, and learning — yes. For confirmatory and published research, verify the results against lme4 (R), MixedModels.jl (Julia), or PROC MIXED (SAS), all of which use restricted maximum likelihood. Cite this tool as: StatsUnlock. (2025). GLMM crossed and nested random effects calculator. https://statsunlock.com.
Q10. My result is non-significant — does that mean the random effect is unimportant?
Not necessarily. A non-significant variance component can mean the effect is genuinely small or that you have too few groups to detect it. Always pair the p-value with the variance component and the ICC. If the ICC is non-trivial but the p-value is large, increase the number of groups, not the within-group sample size.
📚 References
The following references support the statistical methods used in this GLMM crossed and nested random effects calculator, covering variance components, intraclass correlation, and best practices in mixed-effects modeling.
- Bates, D., Mächler, M., Bolker, B., & Walker, S. (2015). Fitting linear mixed-effects models using lme4. Journal of Statistical Software, 67(1), 1–48. https://doi.org/10.18637/jss.v067.i01
- Pinheiro, J. C., & Bates, D. M. (2000). Mixed-effects models in S and S-PLUS. Springer. https://doi.org/10.1007/b98882
- Snijders, T. A. B., & Bosker, R. J. (2012). Multilevel analysis: An introduction to basic and advanced multilevel modeling (2nd ed.). SAGE Publications.
- Goldstein, H. (2011). Multilevel statistical models (4th ed.). Wiley. https://doi.org/10.1002/9780470973394
- Raudenbush, S. W., & Bryk, A. S. (2002). Hierarchical linear models: Applications and data analysis methods (2nd ed.). SAGE Publications.
- Nakagawa, S., & Schielzeth, H. (2013). A general and simple method for obtaining R² from generalized linear mixed-effects models. Methods in Ecology and Evolution, 4(2), 133–142. https://doi.org/10.1111/j.2041-210x.2012.00261.x
- Baayen, R. H., Davidson, D. J., & Bates, D. M. (2008). Mixed-effects modeling with crossed random effects for subjects and items. Journal of Memory and Language, 59(4), 390–412. https://doi.org/10.1016/j.jml.2007.12.005
- Barr, D. J., Levy, R., Scheepers, C., & Tily, H. J. (2013). Random effects structure for confirmatory hypothesis testing: Keep it maximal. Journal of Memory and Language, 68(3), 255–278. https://doi.org/10.1016/j.jml.2012.11.001
- Bolker, B. M., Brooks, M. E., Clark, C. J., Geange, S. W., Poulsen, J. R., Stevens, M. H. H., & White, J. S. (2009). Generalized linear mixed models: A practical guide for ecology and evolution. Trends in Ecology & Evolution, 24(3), 127–135. https://doi.org/10.1016/j.tree.2008.10.008
- Schielzeth, H., & Nakagawa, S. (2013). Nested by design: Model fitting and interpretation in a mixed model era. Methods in Ecology and Evolution, 4(1), 14–24. https://doi.org/10.1111/j.2041-210x.2012.00251.x
- Cohen, J. (1988). Statistical power analysis for the behavioral sciences (2nd ed.). Lawrence Erlbaum Associates.
- American Psychological Association. (2020). Publication manual of the American Psychological Association (7th ed.). APA. https://doi.org/10.1037/0000165-000
- Gelman, A., & Hill, J. (2007). Data analysis using regression and multilevel/hierarchical models. Cambridge University Press. https://doi.org/10.1017/CBO9780511790942
- R Core Team. (2024). R: A language and environment for statistical computing. R Foundation for Statistical Computing. https://www.R-project.org/
- NIST/SEMATECH. (2013). e-Handbook of statistical methods. National Institute of Standards and Technology. https://www.itl.nist.gov/div898/handbook/